Plant-ProChip 2.0: High throughput transformation of plant protoplasts
This project uses microfluidics to develop both transient and stable protoplast transformation protocols on-chip, at a high throughput scale. This project follows on from the following projects:
The results of this project have been published in a publication in PLoS One:
Yu Z, Boehm CR, Hibberd JM, Abell C, Haseloff J, Burgess SJ, Reyna-Llorens I (2018). Droplet-based microfluidic analysis and screening of single plant cells. PLoS One 13(5):e0196810. doi: 10.1371/journal.pone.0196810.
The Idea
A current limitation for plant synthetic biology involves the high-throughput screening of genetic parts in plant cells. Current validation techniques involve fusing sequences to a reporter and analysing expression in planta, which requires testing each circuit in an individual plant, either through transient biolistic transformation or the generation of stable transgenics (Brown et al. 2011). Applying these techniques to whole regulatory element libraries or to a considerable number of variations of a is not feasible at a laboratory scale.
In a previous project funded by the SynbioFund we aimed to develop a high-throughput screen for the analysis of promoter sequences driving the expression of a reporter gene in plant protoplasts. As a result of this project we successfully managed to isolate, encapsulate and analyse protoplasts from the model species, Marchantia polymorpha, Arabidopsis thaliana and Nicotiana benthamiana using a PDMS microfluidic device (Figure 1).
Despite this success, we realised that the efficiency of protoplast transformation now limits the assay, preventing them from being truly high-throughput. For instance, current methods for protoplast transformation using polyethylene glycol (PEG) require high amounts of plasmid, implying many resources need to be invested for the purification of a small number of constructs.
The aim of this project is to make use of microfluidics to develop both transient and stable protoplast transformation protocols on-chip, at a high throughput scale. In this case wild type protoplasts will be isolated and encapsulated with our current methodologies. Protoplasts will be then transformed “in chipo” either by PEG transformation or by co-incubation with Agrobacterium tumefaciens. Droplets carrying the transformed protoplasts will be incubated and screened for reporter activity. Finally, transformed cells will be sorted and plated onto regeneration media with the aim of regenerating whole plants from single transformed protoplasts.
We envisage this system to be applicable to a range of plant species not just for testing DNA parts but to other applications such as the generation of random mutagenesis lines, enhancer trap lines or inserting novel pathways in plants using minimal amount of resources.
The Team
Dr Ivan Reyna-Llorens,
Post-doctoral Researcher, Department of Plant Sciences, University of Cambridge
Dr Steven Burgess,
Post-doctoral Researcher, Department of Plant Sciences, University of Cambridge
Dr Ziyi Yu,
Post-doctoral Researcher, Department of Chremistry, University of Cambridge
Mr Gregory Reeves,
Graduate Researcher, Department of Plant Sciences, University of Cambridge
Mr Christian R. Boehm,
Graduate Researcher, Department of Plant Sciences, University of Cambridge
Project Outputs
Project Report
Summary of the project's achievements and future plans
Project Proposal
Original proposal and application
Project Resources
Optimization of droplet sorting for high throughput transformation of plant protoplasts
Summary
A current limitation for plant synthetic biology involves high-throughput screening of genetic parts in plants. Current approaches require testing circuits in individual plants, through transient or stable transgenics. Applying these techniques to entire libraries is not feasible at a laboratory scale.
In the first stage of the project we aimed to develop a high-throughput screen for the analysis of promoter sequences in plant protoplasts. As a result, we successfully isolated, encapsulated and analysed protoplasts from the model species, Marchantia polymorpha and Arabidopsis thaliana using a PDMS microfluidic device. Despite of this, there are considerable limitations in terms of protoplast transformation for making these assays high-throughput.
The aim of this project is to use microfluidics to develop both transient and stable protoplast transformation protocols at a high-throughput scale. Encapsulated protoplasts will be transformed by PEG transformation and screened for reporter activity. The transformed cells will be sorted and plated onto regeneration media for whole plants regeneration.
We envisage this system to be applicable to a range of plant species not just for testing DNA parts but to other applications such as the generation of random mutagenesis lines, enhancer trap lines or inserting novel pathways in plants using minimal amount of resources.
Report and outcomes
Figure 1: Marchantia polymorpha protoplasts encapsulated in microdroplets.
In the last stage of the project we isolated and encapsulated protoplasts from different species. However, to succeed in transforming protoplast “on chip” we had to optimize the sorting of transformed protoplasts. During the last months, we have been optimizing this aspect of the project as well as fine tuning our method of protoplast isolation.
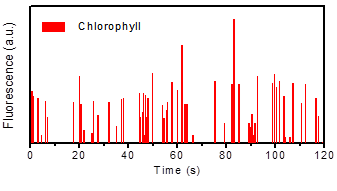
Figure 2: The number of Marchantia protoplasts in microdroplets have been counted by fluorescence-based microdroplets counting system. Each peak represents the chlorophyll fluorescence of one Marchantia protoplast.
We decided to use Marchantia polymorpha protoplasts (Figure 1 & 2) as the model species for this part of the project. In our hands, Marchantia gives the highest yields as well as the maximum protoplast integrity when compared with other systems such as Arabidopsis or tobacco.
Protoplast transformation requires an accurate sorting step in which transformed cells can be selected for further applications such as whole plant regeneration. In this case, we used transgenic Marchantia polymorpha plants carrying the pCRB mpt0 plasmid that contains a constitutive promoter expressing a yellow fluorescent protein (YFP). Protoplasts were isolated from this line and the activity of YFP was assessed (Figure 3).
Figure 3: A) Quantification of YFP in encapsulated marchantia protoplasts. Each peak represents the activity of an individual protoplast. B) Isolated protoplast expressing YFP.
Figure 4: Sorted encapsulated protoplasts from Marchantia polymorpha visualized under A) light microscopy and B) fluorescent microscopy.
We now have an assay to evaluate transgenic (YFP) and a non-transgenic (Chlorophyll) cells that can be used for sorting plant protoplasts. The next step was to sort WT and transgenic Marchantia lines. In a first attempt, we manage to sort chlorophyll positive droplets (Figure 4). However, most of the droplets contained damaged or ruptured protoplasts.
After modifying the isolation of protoplast so most of the cells will retain their integrity, the transgenic lines expressing YFP were encapsulated and sorted using the YFP channel (Figure 5).
Based on these results, we now have a minimal pipeline for protoplast isolation and sorting in Marchantia. In the next phase of the project, we want to assess the activity of different genetic circuits as well as use our chip to optimize protoplast transformation “in chip”.
Figure 5: Marchantia polymorpha protoplasts sorted using a microfluidic device and visualised under A) light microscopy, and a fluorescent microscope using B) a YFP channel (491 nm) and C) a chlorophyll channel (630 nm).